- Home
- About
- Environmental Briefs
-
Distinguished Guest Lectures
- 2023 Water, water, everywhere – is it still safe to drink? The pollution impact on water quality
- 2022 Disposable Attitude: Electronics in the Environment >
- 2019 Radioactive Waste Disposal >
- 2018 Biopollution: Antimicrobial resistance in the environment >
- 2017 Inside the Engine >
- 2016 Geoengineering >
- 2015 Nanomaterials >
- 2014 Plastic debris in the ocean >
- 2013 Rare earths and other scarce metals >
- 2012 Energy, waste and resources >
- 2011 The Nitrogen Cycle – in a fix?
- 2010 Technology and the use of coal
- 2009 The future of water >
- 2008 The Science of Carbon Trading >
- 2007 Environmental chemistry in the Polar Regions >
- 2006 The impact of climate change on air quality >
- 2005 DGL Metals in the environment: estimation, health impacts and toxicology
- 2004 Environmental Chemistry from Space
- Articles, reviews & updates
- Meetings
- Resources
- Index
Antibiotic resistance - from nature to environmental contaminant
Célia M. Manaia
Universidade Católica Portuguesa, CBQF – Centro de Bio-tecnologia e Química Fina – Laboratório Associado, Escola Superior de Bio-tecnologia, Porto, Portugal
[email protected]
ECG Bulletin July 2018
Universidade Católica Portuguesa, CBQF – Centro de Bio-tecnologia e Química Fina – Laboratório Associado, Escola Superior de Bio-tecnologia, Porto, Portugal
[email protected]
ECG Bulletin July 2018
Antibiotic resistance is a human health issue, the consequence of our failure to treat simple bacterial infections. Antibiotic resistance has been disseminated into the environment, and is now a source of biological pollution. Current state-of-the-art research has revealed the most important sources of antibiotic resistance contamination, and provides some suggestions of how it can be controlled.
There are still important uncertainties regarding the human health risks associated with antibiotic resistance. These are due to the complexity of the problem and may be addressed with further research and worldwide surveillance. Until then, it is recommended that the precautionary principle is applied to any activity or source that may contribute to the environmental propagation of antibiotic resistance.
The clinical definition of antibiotic resistance is the failure to treat infections caused by bacteria that were once susceptible to a specific antibiotic or class of antibiotics. Multidrug resistance is due to the accumulation of resistance to different classes of antibiotics in the same bacterial strain. Multidrug resistance leaves, in some cases, very limited therapeutic options. Antibiotic resistance is now a global problem with serious implications for human health and major economic impacts. Worldwide, the number of deaths per year that are attributed to antibiotic resistant bacterial infections is 25,000 in the European Union, 23,000 in the United States of America and 38,000 in Thailand; not only this, but in India in one year more than 58,000 babies died due to infection with resistant bacteria (1). In addition, there are a number of other effects with important economic and health impacts such as extensions to hospitalisation periods and sick leave, and an increase in unsuccessful medical interventions. Antibiotic resistance is not only an issue affecting already diseased persons: it can also affect healthy individuals exposed to antibiotic resistant bacteria, either in healthcare facilities or in any other place where antibiotic resistant bacteria can be found, such as contaminated air or water bodies (2–4).
Although antibiotic resistance is a human health problem handled in healthcare facilities, it did not originate in hospitals. To find the ancestors of clinically relevant antibiotic resistance, it is necessary to investigate the natural environment, where the capacity to specifically resist each class of antibiotics is one of the multiple mechanisms used by bacteria to cope with external stimuli. Therefore, antibiotic resistance somehow escaped from the natural environment, and now exists in human pathogens (5,6). How, why and when this transfer happens is, except in a very few cases, unknown (7, 8).
Over the last 80 years of regular and intensive antibiotic use, antibiotic resistance has been delivered back to the natural environment, and is now a contaminant of great concern. Instead of just being found in strictly environmental bacteria, contaminant antibiotic resistance has become a property of new bacterial hosts, mainly human and animal commensal bacteria and pathogens. These antibiotic resistant bacteria can now be found everywhere, not only in healthy human and animal microbiomes (the whole set of microorganisms inhabiting a given place), but also in the environment, mainly in areas subjected to human activities. The primary sources of antibiotic resistant bacteria are hospital and household sewage and animal production facilities, where antibiotics can be used to promote growth (a practice banned in the European Union in 2006) or to treat infections (9). The impact of these sources on the dissemination of antibiotic resistance varies according to the degree to which good-practices and sanitation are implemented, among other factors (10, 11). Ensuring the availability and sustainable management of water and sanitation is the 6th Sustainable Development Goal proposed by the United Nations (12). In 2015, only 39% of the global population (2.9 billion) used a safely managed sanitation service (13).
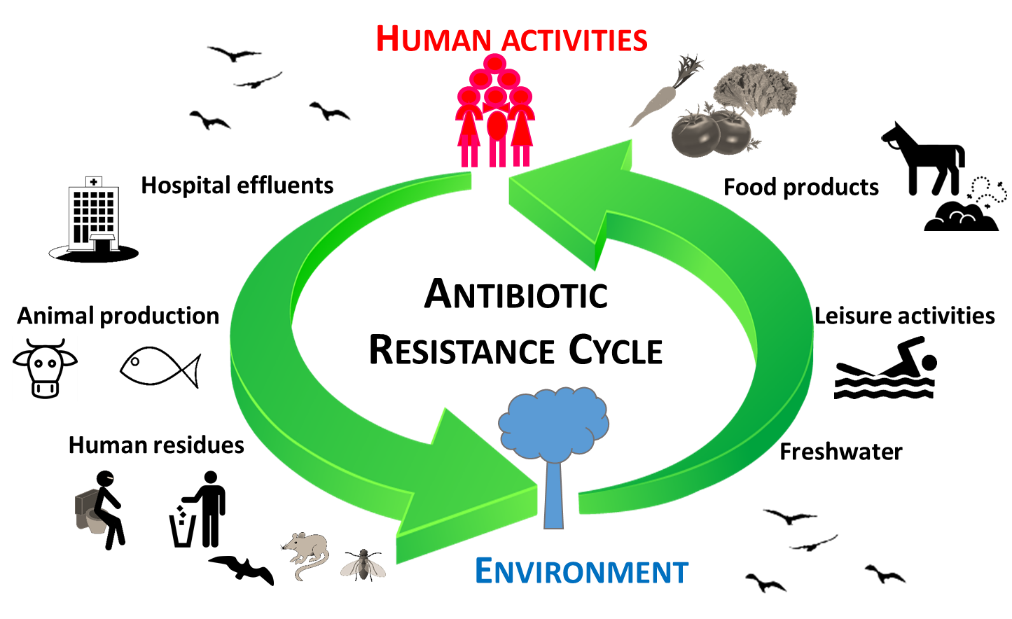
Sewage collection and treatment in waste water treatment plants is unquestionably important for human health and environmental protection. However, multiple studies have shown that state-of-the-art treatment processes are not able to avoid the dissemination of antibiotic resistant bacteria into the environment (14–16). Therefore, even in regions where adequate waste water treatment processes are implemented, the contamination of streams, lakes, rivers, and soils with antibiotic resistant bacteria may be unavoidable (15–18). This problem is aggravated by the fact that, as for many other environmental contaminants, water is a privileged transport route and a preferential bacterial habitat. The generalised distribution of antibiotic resistance in the environment can also result in the contamination of wildlife, the human food chain, or leisure areas (19). The cycle of antibiotic resistance in the environment and the paths of human exposure are extremely complex and still poorly understood (15, 16, 20) (Figure 1).
There are still important uncertainties regarding the human health risks associated with antibiotic resistance. These are due to the complexity of the problem and may be addressed with further research and worldwide surveillance. Until then, it is recommended that the precautionary principle is applied to any activity or source that may contribute to the environmental propagation of antibiotic resistance.
The clinical definition of antibiotic resistance is the failure to treat infections caused by bacteria that were once susceptible to a specific antibiotic or class of antibiotics. Multidrug resistance is due to the accumulation of resistance to different classes of antibiotics in the same bacterial strain. Multidrug resistance leaves, in some cases, very limited therapeutic options. Antibiotic resistance is now a global problem with serious implications for human health and major economic impacts. Worldwide, the number of deaths per year that are attributed to antibiotic resistant bacterial infections is 25,000 in the European Union, 23,000 in the United States of America and 38,000 in Thailand; not only this, but in India in one year more than 58,000 babies died due to infection with resistant bacteria (1). In addition, there are a number of other effects with important economic and health impacts such as extensions to hospitalisation periods and sick leave, and an increase in unsuccessful medical interventions. Antibiotic resistance is not only an issue affecting already diseased persons: it can also affect healthy individuals exposed to antibiotic resistant bacteria, either in healthcare facilities or in any other place where antibiotic resistant bacteria can be found, such as contaminated air or water bodies (2–4).
Although antibiotic resistance is a human health problem handled in healthcare facilities, it did not originate in hospitals. To find the ancestors of clinically relevant antibiotic resistance, it is necessary to investigate the natural environment, where the capacity to specifically resist each class of antibiotics is one of the multiple mechanisms used by bacteria to cope with external stimuli. Therefore, antibiotic resistance somehow escaped from the natural environment, and now exists in human pathogens (5,6). How, why and when this transfer happens is, except in a very few cases, unknown (7, 8).
Over the last 80 years of regular and intensive antibiotic use, antibiotic resistance has been delivered back to the natural environment, and is now a contaminant of great concern. Instead of just being found in strictly environmental bacteria, contaminant antibiotic resistance has become a property of new bacterial hosts, mainly human and animal commensal bacteria and pathogens. These antibiotic resistant bacteria can now be found everywhere, not only in healthy human and animal microbiomes (the whole set of microorganisms inhabiting a given place), but also in the environment, mainly in areas subjected to human activities. The primary sources of antibiotic resistant bacteria are hospital and household sewage and animal production facilities, where antibiotics can be used to promote growth (a practice banned in the European Union in 2006) or to treat infections (9). The impact of these sources on the dissemination of antibiotic resistance varies according to the degree to which good-practices and sanitation are implemented, among other factors (10, 11). Ensuring the availability and sustainable management of water and sanitation is the 6th Sustainable Development Goal proposed by the United Nations (12). In 2015, only 39% of the global population (2.9 billion) used a safely managed sanitation service (13).
Sewage collection and treatment in waste water treatment plants is unquestionably important for human health and environmental protection. However, multiple studies have shown that state-of-the-art treatment processes are not able to avoid the dissemination of antibiotic resistant bacteria into the environment (14–16). Therefore, even in regions where adequate waste water treatment processes are implemented, the contamination of streams, lakes, rivers, and soils with antibiotic resistant bacteria may be unavoidable (15–18). This problem is aggravated by the fact that, as for many other environmental contaminants, water is a privileged transport route and a preferential bacterial habitat. The generalised distribution of antibiotic resistance in the environment can also result in the contamination of wildlife, the human food chain, or leisure areas (19). The cycle of antibiotic resistance in the environment and the paths of human exposure are extremely complex and still poorly understood (15, 16, 20) (Figure 1).
However, the current knowledge in the field paves the way to propose important measures that may contribute to mitigation and control of antibiotic resistance propagation. A first step is the identification of critical antibiotic resistance sources. Two clear examples can be given. Firstly, hospital effluents which may contain high doses of antibiotic resistant bacteria of clinical relevance, and often resistance types that are emerging in the clinical setting but not previously found in the environment. These can be discharged into municipal waste water treatment plants without prior treatment. Clinical antibiotic resistance genes released in hospital effluents can in this way be further disseminated to the environment (18, 21, 22). Secondly, waste water treatment sludge and animal manure, known to be impregnated with antibiotic resistance determinants, can be used as natural fertilisers in soils with serious implications for soil contamination (23, 24). The dedicated treatment of hospital effluents before discharging in the municipal collector, the improvement of municipal waste water treatment to optimise simultaneous removal of antibiotic resistance with the myriad of contaminants that occur in waste water, and the implementation of stewardship measures to minimise antibiotic use in humans and animals are all part of a long list of possible control strategies (10, 14, 16, 24).
Although the scientific community has already identified critical control points (such as those mentioned above), it remains difficult to devise global strategic interventions. It is important to recognise that antibiotic resistance is spread between different environmental compartments, is often present in very high abundance, and represents the most serious clinical threats (e.g. carbapenem and colistin resistance). It is necessary to recognise the paths by which resistance moves from the environment to humans and animals (20). Only some of the wide diversity of antibiotic resistant bacteria in the environment will be able to colonise and eventually infect humans (25). Priorities include the elucidation of important antibiotic resistant vectors that are able to simultaneously thrive in the environment and colonise and/or infect humans. In addition, it is essential to identify the major paths through which antibiotic resistance vectors spread. Another key question refers to the number of cells and associated circumstances that will be needed to cause any harm in the human body. This value corresponds to the designated “infectious dose”, which is not known and is not trivial to determine for most environmental antibiotic resistant bacteria. If known, this value could be used as a reference to determine the maximum admissible load of antibiotic resistant bacteria in a given environment that does not represent a risk to humans. However, the calculation of an “infectious dose” presents other challenges, such as the capacity for bacteria to self-replicate. Another challenge is that the limits of quantification of antibiotic resistance achieved by state-of-the-art techniques, like quantitative PCR, may be too high to enforce reliable environmental safety guidelines. These arguments show that while it may be difficult to provide clear evidence that antibiotic resistance can be transmitted from environmental sources to humans, or to define objective thresholds of maximum admissible values, the application of the precautionary principle to contain the environmental spread and human transmission of antibiotic resistance is urgently needed.
References
1. Centers for Disease Prevention and Control (CDC). (2017). Infographic: Antibiotic Resistance The Global Threat. https://www.cdc.gov/globalhealth/infographics/default.html
2. Vaz-Moreira, I., Nunes, O. C., & Manaia, C. M. (2014). Bacterial diversity and antibiotic resistance in water habitats: searching the links with the human microbiome. FEMS Microbiology Reviews, 38, 761-778.
3. Pal, C. et al. (2016). The structure and diversity of human, animal and environmental resistomes. Microbiome, 4, 54.
4. Bengtsson-Palme, J. et al. (2017). Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, 42, fux053.
5. Allen, H. K. et al. (2010). Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251.
6. Van Goethem, M. W. et al. (2018). A reservoir of ‘historical’antibiotic resistance genes in remote pristine Antarctic soils. Microbiome, 6, 40.
7. D’Andrea, M. M. et al. (2013). CTX-M-type β-lactamases: a successful story of antibiotic resistance. International Journal of Medical Microbiology, 303(6-7), 305-317.
8. Nordmann, P., & Poirel, L. (2005). Emergence of plasmid-mediated resistance to quinolones in Enterobacteriaceae. Journal of Antimicrobial Chemotherapy, 56, 463-469.
9. Davies, J., & Davies, D. (2010). Origins and evolution of antibiotic resistance. Microbiology and Molecular Biology Reviews, 74, 417-433.
10. Berendonk, T. U. et al. (2015). Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310.
11. Wuijts, S. et al. (2017). Towards a research agenda for water, sanitation and antimicrobial resistance. Journal of Water and Health, 15, 175-184.
12. United Nations (UN). Transforming our world: the 2030 agenda for sustainable development. (2015) A/RES/70/1. sustainabledevelopment.un.org
13. World Health Organization (WHO) (2015). News Rooms – facts sheet. Sanitation.
14. Bürgmann, H, et al. (2018). Water & Sanitation: An Essential Battlefront in the War on Antimicrobial Resistance FEMS Microbiology Ecology (In press)
15. Manaia, C. M. et al. (2016). Antibiotic resistance in urban aquatic environments: can it be controlled?. Applied Microbiology and Biotechnology, 100, 1543-1557.
16. Vikesland, P. J. et al. (2017). Toward a comprehensive strategy to mitigate dissemination of environmental sources of antibiotic resistance.
17. Czekalski, N., Díez, E. G., & Bürgmann, H. (2014). Wastewater as a point source of antibiotic-resistance genes in the sediment of a freshwater lake. The ISME Journal, 8, 1381.
18. Vaz-Moreira, I. et al. (2016). Multidrug resistance in quinolone-resistant gram-negative bacteria isolated from hospital effluent and the municipal wastewater treatment plant. Microbial Drug Resistance, 22, 155-163.
19. Marti, E. et al. (2018). Abundance of antibiotic resistance genes and bacterial community composition in wild freshwater fish species. Chemosphere, 196, 115-119.
20. Larsson, D.G.J. et al. (2018). Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environment International (in press).
21. Narciso-da-Rocha, C. et al. (2014). blaTEM and vanA as indicator genes of antibiotic resistance contamination in a hospital–urban wastewater treatment plant system. Journal of Global Antimicrobial Resistance, 2, 309-315.
22. Varela, A. R. et al. (2015). Molecular evidence of the close relatedness of clinical, gull and wastewater isolates of quinolone-resistant Escherichia coli. Journal of Global Antimicrobial Resistance, 3, 286-289.
23. McKinney, C. W. et al. (2018). Occurrence and abundance of antibiotic resistance genes in agricultural soil receiving dairy manure. FEMS Microbiology Ecology.
24. Topp E. et al. (2018). Antimicrobial resistance and the environment: assessment of advances, gaps and recommendations for agriculture, aquaculture and pharmaceutical manufacturing. FEMS Microbiology Ecology, 1, 94. doi: 10.1093/femsec/fix185.
25. Manaia, C. M. (2017). Assessing the risk of antibiotic resistance transmission from the environment to humans: non-direct proportionality between abundance and risk. Trends in Microbiology, 25, 173-181.
Although the scientific community has already identified critical control points (such as those mentioned above), it remains difficult to devise global strategic interventions. It is important to recognise that antibiotic resistance is spread between different environmental compartments, is often present in very high abundance, and represents the most serious clinical threats (e.g. carbapenem and colistin resistance). It is necessary to recognise the paths by which resistance moves from the environment to humans and animals (20). Only some of the wide diversity of antibiotic resistant bacteria in the environment will be able to colonise and eventually infect humans (25). Priorities include the elucidation of important antibiotic resistant vectors that are able to simultaneously thrive in the environment and colonise and/or infect humans. In addition, it is essential to identify the major paths through which antibiotic resistance vectors spread. Another key question refers to the number of cells and associated circumstances that will be needed to cause any harm in the human body. This value corresponds to the designated “infectious dose”, which is not known and is not trivial to determine for most environmental antibiotic resistant bacteria. If known, this value could be used as a reference to determine the maximum admissible load of antibiotic resistant bacteria in a given environment that does not represent a risk to humans. However, the calculation of an “infectious dose” presents other challenges, such as the capacity for bacteria to self-replicate. Another challenge is that the limits of quantification of antibiotic resistance achieved by state-of-the-art techniques, like quantitative PCR, may be too high to enforce reliable environmental safety guidelines. These arguments show that while it may be difficult to provide clear evidence that antibiotic resistance can be transmitted from environmental sources to humans, or to define objective thresholds of maximum admissible values, the application of the precautionary principle to contain the environmental spread and human transmission of antibiotic resistance is urgently needed.
References
1. Centers for Disease Prevention and Control (CDC). (2017). Infographic: Antibiotic Resistance The Global Threat. https://www.cdc.gov/globalhealth/infographics/default.html
2. Vaz-Moreira, I., Nunes, O. C., & Manaia, C. M. (2014). Bacterial diversity and antibiotic resistance in water habitats: searching the links with the human microbiome. FEMS Microbiology Reviews, 38, 761-778.
3. Pal, C. et al. (2016). The structure and diversity of human, animal and environmental resistomes. Microbiome, 4, 54.
4. Bengtsson-Palme, J. et al. (2017). Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, 42, fux053.
5. Allen, H. K. et al. (2010). Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251.
6. Van Goethem, M. W. et al. (2018). A reservoir of ‘historical’antibiotic resistance genes in remote pristine Antarctic soils. Microbiome, 6, 40.
7. D’Andrea, M. M. et al. (2013). CTX-M-type β-lactamases: a successful story of antibiotic resistance. International Journal of Medical Microbiology, 303(6-7), 305-317.
8. Nordmann, P., & Poirel, L. (2005). Emergence of plasmid-mediated resistance to quinolones in Enterobacteriaceae. Journal of Antimicrobial Chemotherapy, 56, 463-469.
9. Davies, J., & Davies, D. (2010). Origins and evolution of antibiotic resistance. Microbiology and Molecular Biology Reviews, 74, 417-433.
10. Berendonk, T. U. et al. (2015). Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310.
11. Wuijts, S. et al. (2017). Towards a research agenda for water, sanitation and antimicrobial resistance. Journal of Water and Health, 15, 175-184.
12. United Nations (UN). Transforming our world: the 2030 agenda for sustainable development. (2015) A/RES/70/1. sustainabledevelopment.un.org
13. World Health Organization (WHO) (2015). News Rooms – facts sheet. Sanitation.
14. Bürgmann, H, et al. (2018). Water & Sanitation: An Essential Battlefront in the War on Antimicrobial Resistance FEMS Microbiology Ecology (In press)
15. Manaia, C. M. et al. (2016). Antibiotic resistance in urban aquatic environments: can it be controlled?. Applied Microbiology and Biotechnology, 100, 1543-1557.
16. Vikesland, P. J. et al. (2017). Toward a comprehensive strategy to mitigate dissemination of environmental sources of antibiotic resistance.
17. Czekalski, N., Díez, E. G., & Bürgmann, H. (2014). Wastewater as a point source of antibiotic-resistance genes in the sediment of a freshwater lake. The ISME Journal, 8, 1381.
18. Vaz-Moreira, I. et al. (2016). Multidrug resistance in quinolone-resistant gram-negative bacteria isolated from hospital effluent and the municipal wastewater treatment plant. Microbial Drug Resistance, 22, 155-163.
19. Marti, E. et al. (2018). Abundance of antibiotic resistance genes and bacterial community composition in wild freshwater fish species. Chemosphere, 196, 115-119.
20. Larsson, D.G.J. et al. (2018). Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environment International (in press).
21. Narciso-da-Rocha, C. et al. (2014). blaTEM and vanA as indicator genes of antibiotic resistance contamination in a hospital–urban wastewater treatment plant system. Journal of Global Antimicrobial Resistance, 2, 309-315.
22. Varela, A. R. et al. (2015). Molecular evidence of the close relatedness of clinical, gull and wastewater isolates of quinolone-resistant Escherichia coli. Journal of Global Antimicrobial Resistance, 3, 286-289.
23. McKinney, C. W. et al. (2018). Occurrence and abundance of antibiotic resistance genes in agricultural soil receiving dairy manure. FEMS Microbiology Ecology.
24. Topp E. et al. (2018). Antimicrobial resistance and the environment: assessment of advances, gaps and recommendations for agriculture, aquaculture and pharmaceutical manufacturing. FEMS Microbiology Ecology, 1, 94. doi: 10.1093/femsec/fix185.
25. Manaia, C. M. (2017). Assessing the risk of antibiotic resistance transmission from the environment to humans: non-direct proportionality between abundance and risk. Trends in Microbiology, 25, 173-181.